
R E S E A R C H
Open Access
Long non-coding RNA LINC00520 promotes
the proliferation and metastasis of
malignant melanoma by inducing the miR-
125b-5p/EIF5A2 axis
Wenkang Luan
1*
†
, Yuting Ding
2
†
, Haitao Yuan
3
†
, Shaojun Ma
1
, Hongru Ruan
1
, Jinlong Wang
1
, Feng Lu
1
and
Xuefeng Bu
3*
Abstract
Background:
Long intergenic non-protein coding RNA 520 (LINC00520), a novel identified lncRNA, has been
shown to modulate the malignant phenotype of tumor cells in some malignant tumors. However, the exact role
and molecular mechanism of LINC00520 in malignant melanoma has not been studied.
Methods:
The expression of LINC00520 in melanoma tissues were detected by using RNA-seq analysis and qRT-
PCR. Melanoma cases from the public databases (The Cancer Genome Atlas (TCGA), GEO#GSE15605, GEO#GSE34460
and GEO#GSE24996) were included in this study. CCK-8 assay, EdU assay, transwell and scratch wound assay were
used to explore the role of LINC00520 in melanoma cells. Luciferase reporter assays, MS2-RIP, RNA pull-down and
RNA-ChIP assay were used to demonstrate the molecular biological mechanism of LINC00520 in melanoma.
Results:
We found that LICN00520 was found to be overexpressed in melanoma tissue. High expression of
LICN00520 is a risk factor for the prognosis of melanoma patients. LINC00520 promotes the proliferation, invasion
and migration of melanoma cells. LICN00520 exerted its oncogenic role by competitive binding miR-125b-5p to
promote Eukaryotic initiation factor 5A2 (EIF5A2) expression. We also showed that LICN00520 promotes the growth
and metastasis of melanoma in vivo through regulating miR-125b-5p/EIF5A2 axis.
Conclusions:
All results elucidated the role and molecular mechanism of LINC00520 in the malignant development
of melanoma. LINC00520, a new oncogene in melanoma, maybe serve as a survival biomarkers or therapeutic
target for melanoma patients.
Keywords:
Melanoma, Growth and metastasis, LICN00520, miR-125b-5p, EIF5A2
© The Author(s). 2020
Open Access
This article is licensed under a Creative Commons Attribution 4.0 International License,
which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give
appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if
changes were made. The images or other third party material in this article are included in the article's Creative Commons
licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons
licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain
permission directly from the copyright holder. To view a copy of this licence, visit
.
The Creative Commons Public Domain Dedication waiver
applies to the
data made available in this article, unless otherwise stated in a credit line to the data.
* Correspondence:
†
Wenkang Luan, Yuting Ding and Haitao Yuan contributed equally to this
work.
1
Department of Plastic Surgery, Affiliated People
’
s Hospital of Jiangsu
University, 8 Dianli Road, Zhenjiang 212000, Jiangsu, China
3
Department of General Surgery, Affiliated People
’
s Hospital of Jiangsu
University, 8 Dianli Road, Zhenjiang 212000, Jiangsu, China
Full list of author information is available at the end of the article
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
https://doi.org/10.1186/s13046-020-01599-7

Introduction
Malignant melanoma is the most dangerous skin tumor,
which is the primary cause of death of skin cancer
In recent years, the global incidence of melanoma is
growing rapidly each year
Although melanoma
patients are treated by the combination of surgery,
chemotherapy, targeted therapy and immunotherapy,
the therapeutic effect is still unsatisfactory, especially
those with distant metastasis [
]. The development of
melanoma is closely related to the abnormal regulation
of multiple genes and signaling pathways
Therefore,
it is very important to explore the molecular mechanism
of the malignant progression of melanoma and to find
potential therapeutic targets for melanoma.
Long non-coding RNAs (lncRNAs), a kind of non-coding
RNA over 200 nucleotides in length, play pivotal roles in
various human tumors by modulating the malignant pheno-
type of tumor cells
A few lncRNAs has been found to
be abnormally expressed in melanoma and involved in its
malignant progress
]. Long intergenic non-protein
coding RNA 520 (LINC00520), located on chromosome 14,
has been reported to overexpress and function as a onco-
gene in breast cancer, nasopharyngeal carcinoma and laryn-
geal squamous cell carcinoma
It has also been
showed that LINC00520 inhibits the growth and metastasis
of cutaneous squamous cell carcinoma
However, the
exact role and molecular mechanism of LINC00520 in
malignant melanoma has not been studied.
In the present study, we found that LINC00520 was
highly expressed in melanoma by analyzing the lncRNAs
expression profile of melanoma. LINC00520 promoted the
proliferation and metastasis of melanoma. To date, the
most widely studied mechanism of lncRNAs in tumor is
that lncRNAs play the role of competitive endogenous
RNAs (ceRNA) in tumor development
Similarly,
LINC00520 has been shown to play the same role in naso-
pharyngeal carcinoma [
. Here, we have established the
ceRNA regulatory network of LINC00520 based on RNA-
seq and miRNA-seq results and bioinformatics predictions.
We demonstrated that LINC00520 exerts its oncogene ef-
fect in melanoma by regulating Eukaryotic initiation factor
5A2 (EIF5A2). EIF5A2, SUPPLlocated on human chromo-
some 3q25
–
27, function as a novel oncogene in many tu-
mors
]. Extensive studies have demonstrated that
EIF5A2 participates in the proliferation, migration, invasion
and chemotherapeutic resistance of hepatocellular carcin-
oma, esophageal cancer, gastric cancer, melanoma, etc.
[
]. We proved that miR-125b-5p exerts anti-cancer
effects in melanoma by targeting EIF5A2. Furthermore, we
showed that LICN00520 can remove the inhibition effect of
miR-125b-5p on EIF5A2 through decoying miR-125b-5p,
thus promoting the expression of EIF5A2. Therefore,
LICN00520 can serve as a new special diagnostic indicator
and therapeutic target in melanoma patients.
Materials and methods
Tissue samples
Forty-one primary malignant melanoma tissues and adja-
cent normal tissues (ANT) were collected from the melan-
oma patients in the Affiliated People
’
s Hospital of Jiangsu
University, and informed consent was obtained from all pa-
tients.. The pathological diagnosis was made independently
by two pathologists. None of the patients had undergone
chemotherapy or radiotherapy. The study was approved by
the Human Research Ethics Committee of the Affiliated
People
’
s Hospital of Jiangsu University. The public database
of melanoma from The Cancer Genome Atlas (TCGA),
GEO#GSE15605, GEO#GSE34460 and GEO#GSE24996
were also included in this study.
RNA-seq, miRNA-seq and ceRNA analysis
Three melanoma tissues and adjacent normal tissues
were stored in liquid nitrogen, and TRIzol (Invitrogen,
USA) was used to extract RNA. Gminix (Shanghai,
China) conducted the RNA-seq and miRNA-seq ana-
lysis. The network of LICN00520-miRNA-target gene
was constructed by using Cytoscape software (v.3.6.0)
based on the RNA-seq and miRNA-seq results. The
interaction between LINC00520 and miRNAs was pre-
dicted through miRcode. TargetScan, miRDB and miR-
TarBase were used to find the target genes of miRNAs.
Cell lines and cell culture
Human malignant melanoma cell lines (A375, A2058,
MeWo, CHL-1, SK-MEL-28) were obtained from the
American Type Culture Collection (ATCC, USA), and
growed in Dulbecco
’
s modified Eagle
’
s medium (DMEM;
Gibco, USA) with 10% fetal bovine serum (Invitrogen,
USA). Human epidermal melanocytes (HEMa-LP) was
purchased from Invitrogen (USA), and maintained in
medium 254 (Cascade Biologics, USA). These cell lines
were incubated in the humidified incubator with the
atmosphere of 37 °C containing 5% CO
2
.
Plasmids, oligonucleotides and transfection
The miR-125b-5p mimic, miR-125b-5p inhibitor and re-
lated negative control were obtained by GenePharma
(Shanghai, China). The small interfering RNA (siRNA)
and short hairpin RNA (shRNA) of LINC00520 were
also chemically synthesized by GenePharma (Shanghai,
China). The EIF5A2 plasmid was constructed by insert-
ing the full length of EIF5A2 into pcDNA3.1 vector
(Invitrogen, USA). The shRNA and the control were
inserted into the lentivirus vector (GenePharma, Shang-
hai, China), and the stably expressing sh-LINC00520
shRNA A375 cells were constructed by infecting cells
with the corresponding lentivirus. Lipofectamine 3000
(Invitrogen, USA) was used to transfect the related oli-
gonucleotides into melanoma cells.
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 2 of 16

Quantitative RT-PCR
TRIzol reagent was used to extract RNA from cells and
tissues according to the specified steps (Invitrogen,
USA). Fermentas and microRNA reverse transcription
kits (Applied Biosystems, CA) were used to conduct reverse
transcription. The amplification reactions were conducted
by using the ABI StepOnePlus System (Applied Biosystems,
CA) according to the set reaction conditions. The special
primer of miR-125b-5p was purchased from RiboBio
(Guangzhou, China). GAPDH and U6 was used for
normalization respectively. The following primers were
used:
LINC00520
forward
5
′
-CCTGCTCCTTCAGG
GACATC-3
′
and LINC00520 reverse 5
′
-TCCGCCCCTT
GCTCAAATAG-3
′
; EIF5A2 forward 5
′
-TTCCAGCACT
TACCCTT-3
′
and EIF5A2 reverse 5
′
-TTCCCCTCTA
TTTTTG-3
′
; GAPDH forward 5
′
- GTCAACGGATTTGG
TCTGTATT-3
′
and GAPDH reverse 5
′
- AGTCTTCTGG
GTGGCAGTGAT-3
′
. The method of 2
–
△△
Ct
was used to
calculate the relative expression level.
Western blot
RIPA buffer (KenGEN, China) was used to extract the
protein following the appropriate steps. BCA Protein
Assay Kit (Beyotime, China) was used to measure the con-
centration of extracted protein. Western blotting is carried
out as the previous described
Antibodies against
EIF4A2 (Abcam, 1:1000, Cambridgeshire, UK), vimentin
(Abcam,
1:2000,
Cambridgeshire,
UK),
E-cadherin
(Abcam, 1:500, Cambridgeshire, UK), N-cadherin (Abcam,
1:1000, Cambridgeshire, UK) was used to the related pro-
tein level.
β
-actin (1:1000, Abcam, UK) and GAPDH (1:
2500, Abcam, UK) were used for normalization.
Cell proliferation assay
For cell counting kit-8 (CCK-8, Beyotime, Shanghai,
China) assay, the transfected melanoma cells (5000 cells)
were seeded in a 96-well plate, and the process is carried
out as described previously [
Microplate reader
(Multiscan FC, Thermo Scientific) was used to measure
the absorbance at an optical density of 450 nm. For EdU
assay, the DNA synthesis of melanoma cells grown was
measured by using a EdU imaging kit (life Technologies,
USA). The assay were carried out according to the man-
ufacturer
’
s instructions. Immunostaining were visualized
by using Leica DMI3000B microscope, and the positive
cells were counted.
Cell invasion and migration assays
Transwell assay was used to detect the invasiveness of
melanoma
cell.
Transfected
melanoma
cells
were
digested and resuspended in serum-free DMEM, and
were placed at the top of the Matrigel-coated chambers
(BD Biosciences, USA). The culture medium with 10%
fetal bovine serum was used as the chemical attractant
and added to the lower chamber. After 24 h, the fixed
invasive cells were stained with crystal violet, counted
and photographed. Scratch wound assay was used to
evaluate the migration of melanoma cells. Transfected
melanoma cells were added into the 6-well plates, and
the wound space was formed by the tip of a 200
μ
l pip-
ette. The width of wound was recorded at 0 and 24 h
respectively.
Isolation of RISC-associated RNA
We used 1% formaldehyde to fix miR-125b-5p overexpressed
melanoma cells. We did the chromatin fragmentation.
NETN buffer was used to dissolve the cells, the cells were
then cultured with Dynabeads protein A (Invitrogen, USA)
plus IgG or anti-Pan-Ago, clone 2A8 antibody (Millipore,
USA). We used proteinase K digestion to release immuno-
precipitated RNA. The extracted RNA was purified by glyco-
gen ethanol precipitation and treated with DNase I.
Luciferase reporter assay
The fragment of EIF4A2 3
′
-UTR and LINC00520 con-
taining the miR-125b-5p binding site were inserted into
pMIR-REPORT plasmid, and the mutated plasmid the
used as the control. The corresponding oligonucleotides
and luciferase reporter plasmids were co-transfected into
melanoma cells. The luciferase activity of luciferase
reporter plasmids was measured by Dual Luciferase Re-
porter Assay System (Promega, USA) .
Fluorescence in situ hybridization (FISH)
RiboTM Fluorescent In Situ Hybridization Kit (RiboBio,
Guangzhou, China) was used to for FISH. The proced-
ure was carried out according to the previous study
The probe of LINC00520 was synthesized by RiboBio
(Guangzhou, China). and the cell nucleus were stained
with DAPI. Representative images were obtained by
using a confocal microscopy, and the image J software
was used to collect signals.
MS2-RIP assay
Maltose-binding protein (MBP)-affinity purification was
used to detect miRNAs that binding to LINC00520.
According to the Steitz laboratory steps, MS2-MBP was
purified from E. colicoli. 3 bacteriophage MS2 coat protein
binding sites were inserted in the downstream of
LINC00520 by using Stratagene Quik Change Site Directed
Mutagenesis Kit. The MS2-tagged LINC00520 was trans-
fected into the melanoma cell to obtain miRNAs that asso-
ciated with LINC00520. The RIP analysis was performed
on the cells as previously described after 48 h
], and the
miR-125b-5p level was detected by qRT-PCR.
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 3 of 16

RNA pull-down assay
The Biotinylated of miR-125b-5p was chemical synthe-
sized by GenePharma (Shanghai, China), and the bio-
tinylated mutant and NC were used as control. The
related oligonucleotides were transfected into melanoma
cells. The lysates of cells were cultivated with M-280
streptavidin magnetic beads (Invitrogen, USA)
QRT-PCR was used to detect the LINC00520 level in
the bound RNA.
Xenograft tumor and in vivo lung metastasis assay
10 nude mice were obtained from the Beijing Laboratory
Animal Center (Beijing, China), and these mice were
subcutaneously injected with A375 cells stably express-
ing LINC00520 siRNA. The volume of tumour was mea-
sured every 4 days according to the formula (0.5 × length
× width
2
). After 28 days, mice were sacrificed, and
tumour tissues were stripped and weighed. We injected
A375 cells stably expressing sh-LINC00520 into the tail
vein of mice. 10ul/g sterile D-Luciferin firefly potassium
salt (Beyotime, China) were intraperitoneal injected into
8 nude mice, and the PerkinElmer IVIS Spectrum
(Xenogen, CA) was used for in vivo imaging. The results
were quantified by using the Living Image software
(Xenogen, CA). After 20 days, the lung was dissected
and the metastatic nodules were counted. The study was
approved by the Experimental Animal Ethics Committee
of the Affiliated People
’
s Hospital of Jiangsu University.
Immunohistochemistry staining and HE staining
Immunohistochemistry is performed as described previ-
ously [
using the antibody EIF4A2. The optical density
of the image was analyzed by image J software. For HE
staining, the sections were deparaffinizated and rehydrated.
Then, the sections were incubated with hematoxylin and
stained in acid ethanol and eosin. The sections were dehy-
drated with alcohol and cleared with xylene. Representative
images were taken with a microscope.
Statistical analysis
Data expressed as mean ± SD. SPSS13.0 was used to ana-
lyse the data. Data was evaluated by t-test or one-way
ANOVA, and spearman correlation analysis was ana-
lysed by using the MATLAB. Kaplan-Meier survival
curves was used to evaluate the relationship between
LINC00520 expression and melanoma patient survival.
Melanoma tissues were separated into two groups ac-
cording to the expression of LINC00520, the differences
between the curves were tested by the log-rank test.
GraphPad Prism was used to plot the Kaplan-Meier sur-
vival curves.
P
value < 0.05 is statistically significant.
Results
LINC00520 was significantly up-regulated in melanoma
We first analysed the lncRNA expression profiling in three
malignant melanoma tissues and three adjacent normal tis-
sues (ANT) by using RNA-seq. Volcano plots showed the
differentially expressed lncRNAs over 2.0-fold change be-
tween melanoma tissues and ANT (Fig.
All lncRNAs
whose expression changes over 3.0-fold were shown in a
cluster heat map (Fig.
b). Thereinto, the LINC00520 level
was up-regulated 7.19-fold in melanoma tissues (Fig.
We verified the RNA-seq results by detecting the expres-
sion of LINC00520 in 38 melanoma tissues and ANT, and
found that LINC00520 was increased in melanoma tissues
(Fig.
c). We also found the same result by analyzing the
published datasets (GEO#GSE15605) (Fig.
. GEPIA
(
was used to analyze the ex-
pression of LINC00520 in melanoma dataset of The Can-
cer Genome Atlas (TCGA), and found that LINC00520
was overexpressed in melanoma (Fig.
). Meanwhile, the
level of LINC00520 in malignant melanoma cell (A375,
A2058, CHL-1, MeWo, SK-MEL-28) was higher than that
in human epidermal melanocytes (HEMa-LP) (Fig.
f).
These suggested that LINC00520 maybe participate in the
malignant development of melanoma.
LINC00520 is an risk factor for the survival of patients
with melanoma
We investigated the clinical significance of LINC00520 in
melanoma patients. The high expression of LINC00520
(expression ratio
≥
median ratio) is closely related to the
clinical stage of melanoma, but not to age, sex, ulcer and
family history (Table
). In our melanoma patient samples,
Kaplan-Meier analysis showed that the survival rate of
melanoma patients with high LINC00520 levels was
poorer (Fig.
a). We next analyzed the melanoma patients
prognostic data of TCGA by using GEPIA
) and Starbase
),
and found that the high LINC00520 levels were correlated
with poor survival rate of melanoma patients (Fig.
b and
c). These demonstrated that high LINC00520 expression
is an risk factor for the melanoma patients.
LINC00520 promotes the proliferation, invasion and
migration of melanoma cell
To explore the influence of LINC00520 on the biological
role of melanoma cell, the LINC00520 siRNA was trans-
fected into A375 and A2058 cells (Fig.
CCK-8 assays
revealed that reduction of LINC00520 significantly re-
pressed the proliferation ability of A375 and A2058 cells
(Fig.
EdU assay showed that the number of EdU-
positive cells in LINC00520 knockdown melanoma cells
were significantly reduced compared with the control
group (Fig.
Increasing evidence show that epithelial-
to-mesenchymal transition (EMT) is a key event in the
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 4 of 16
process of tumor metastasis
]. In EMT, there are
morphological changes epithelial-like to mesenchymal-
like appearance
]. We explored the effects of
LINC00520 on the EMT of melanoma cells. The level of
epithelial
cell
marker
(E-cadherin)
was
increased,
whereas the levels of the mesenchymal markers (N-cad-
herin and vimentin) were decreased in LINC00520
knockdown melanoma cells (Fig.
Transwell assays
demonstrated that LINC00520 siRNA inhibited the inva-
sive capacity of A375 and A2058 cells (Fig.
e). Scratch
wound assays revealed that the migrative capacity of
melanoma cells was suppressed by the LINC00520
siRNA (Fig.
In addition, A375 and A2058 are BRAF
mutated melanoma cells. We repeated cell proliferation,
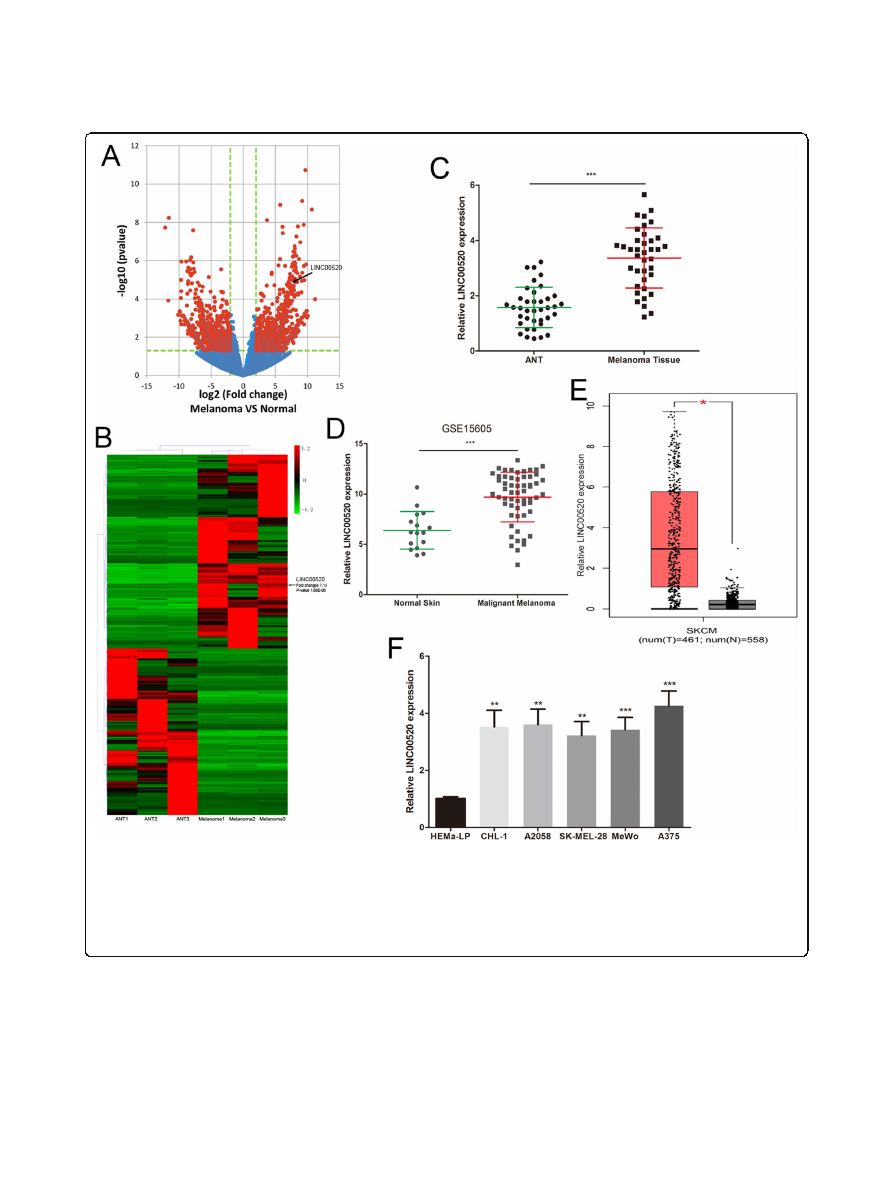
Fig. 1
LINC00520 was significantly up-regulated in melanoma.
a
The volcano plot showed the levels of lncRNAs between primary malignant
melanoma tissues and ANT. The vertical lines represent 2.0-fold changes, and the horizontal line represents
P
-value of 0.05. The red dot
correspond to the differentially expressed lncRNAs with statistical significance.
b
The cluster heat map showed differentially expressed lncRNAs
over 3.0-fold change in melanoma tissues.
c
The level of LINC00520 was analyzed in 38 malignant melanoma tissues and ANT.
d
The LINC00520
levels were detected in the GEO#GSE15605 dataset.
e
GEPIA (
was used to detect the expression of LINC00520 in
TCGA melanoma dataset.
f
The expression profile of LINC00520 in human melanoma cell lines (A375, A2058, MeWo, CHL-1, SK-MEL-28) and
human epidermal melanocytes (HEMa-LP). Data were expressed as the mean ± SD, *
P
< 0.05, **
P
< 0.01, ***
P
< 0.001
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 5 of 16

invasion and migration experiments with BRAF-WT
MeWo cells and reached the same conclusion (Supple-
mentary Fig.
). This suggested that the role of
LINC00520 in melanoma cells is independent of BRAF
mutation.
ceRNA analysis of LINC00520
We
next
explored
the
molecular
mechanism
of
LINC00520 in melanoma. LINC00520 has been shown
to play its role in nasopharyngeal carcinoma through the
ceRNA mechanism [
]. We then constructed the
LINC00520-miRNA-target gene network based on our
miRNA-seq and RNA-seq data (Fig.
The interaction
between LINC00520 and miRNAs was identified using
miRcode, and the target genes of the miRNAs were pre-
dicted through TargetScan, miRDB and miRTarBase.
Cytoscape was used to visualize the interrelationships of
LINC00520-miRNA-target gene. In the network, the
miR-125b-5p/EIF5A2 axis caught our attention because
of miR-125b-5p was decreased and EIF5A2 was in-
creased in melanoma tissue (Fig.
In order to further
verify the results of ceRNA analysis, the expression of
miR-125b-5p and EIF5A2 were detected in 38 melanoma
tissues and ANT. The level of miR-125b-5p was down-
regulated in melanoma tissues (Fig.
b), and the same
result were obtained by analysing the previously pub-
lished dataset (GEO#GSE34460 and GEO#GSE24996)
(Fig.
cand d). EIF5A2 was increased in melanoma tis-
sues compared to ANT (Fig.
e), and the same result
were discovered by analyzing the TCGA-melanoma
dataset (Fig.
f). In our 38 melanoma tissue samples,
LINC00520 and miR-125b-5p levels were inversely cor-
related, while LINC00520 and EIF5A2 levels were
positively correlated (Fig.
and h). Moreover, the
TCGA-melanoma dataset also reveals the same correl-
ation analysis results (Fig.
and j).
LINC00520 sponges miR-125b-5p in melanoma cells
We further explored whether LINC00520 can directly
binds to miR-125b-5p. LINC00520-FISH and qRT-PCR
of nucleus and cytoplasm fragments showed that
LINC00520 was distributed in both cytoplasm and nu-
cleus in melanoma cells (Fig.
a and b). We constructed
the LINC00520 luciferase reporter plasmids containing
the miR-125b-5p binding sites, and the mutated plasmid
was used as the control. (Fig.
The luciferase activity
of wild-type LINC00520 vector was significantly inhib-
ited by the miR-125b-5p mimic in melanoma cells, but
not the mutant plasmid (Fig.
We subsequently verify
the direct binding interaction between LINC00520 and
miR-125b-5p using MS2-RIP and RNA pull-down assay.
The MS2-tagged wild-type LINC00520 vector enriched
lots of miR-125b-5p compared with the empty and mu-
tant plasmids (Fig.
e). Additionally, RNA pull-down
assay also revealed that LINC00520 was pulled down by
biotin-labelled miR-125b-5p (Fig.
The level of miR-
125b-5p was increased in LINC00520 knockdown A375
and A2058 cells (Fig.
All results suggested that
LINC00520 directly binds to miR-125b-5p in melanoma.
LINC00520 acts as a ceRNA to promote EIF5A2 expression
We further discussed the role and mechanism of
LINC00520 on EIF5A2 expression. The 3
′
-UTR of
EIF5A2 has the same miR-125b-5p binding sites that
miR-125b-5p binds to LINC00520 (Fig.
a). We con-
structed the wild-type and mutant EIF5A2 3
’
UTR
Table 1
Correlation between LINC00520 levels and clinical pathological characteristic (
n
= 38)
Clinical characteristics
Number
High LINC00518 expression
Low LINC00518 expression
P
-value
Age
0.494
<50
13
5
8
≥
50
25
14
11
Gender
0.742
Male
22
12
10
Female
16
7
9
Family history
0.403
Yes
7
2
5
No
31
17
14
Ulcer
0.330
Yes
20
12
8
No
18
7
11
TMN stage
<0.01
I-II
14
2
12
III
24
17
7
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 6 of 16
luciferase plasmids, and found that the luciferase activity
of wild type plasmid was suppressed by miR-125b-5p
mimic (Fig.
b). miRNA transfer into Ago2 protein to
form a Ago2/RNA-induced silencing complex (RISC)
[
miRNAs target the binding of RISC to specific
mRNA, leading to mRNA silencing or destabilizing
RNA-ChIP analysis was used to detect the EIF5A2
mRNA abundance in the Ago2/RISC after up-regulation
of miR-125b-5p. miR-125b-5p overexpression melanoma
cells showed the enrichment of the miR-125b-5p and
EIF5A2 level that incorporated into RISC (Fig.
c).
Meanwhile, miR-125b-5p inhibited the expression of
EIF5A2 in both mRNA and protein level. (Fig.
and f).
These demonstrated that EIF5A2 is the target gene of
miR-125b-5p. Furthermore, LINC00520 siRNA also led
a significantly decrease in the luciferase activity of wild
type EIF5A2 3
’
UTR luciferase plasmid, and this effect
could be attenuated by miR-125b-5p inhibitor (Fig.
d).
The mRNA and protein levels of EIF5A2 were repressed
by LINC00520 siRNA in melanoma cells, and this inhib-
ition effect could be reversed by miR-125b-5p inhibitor
(Fig.
e and f). Take together, these results demonstrated
that LINC00520 promotes EIF5A2 expression by spon-
ging miR-125b-5p in melanoma.
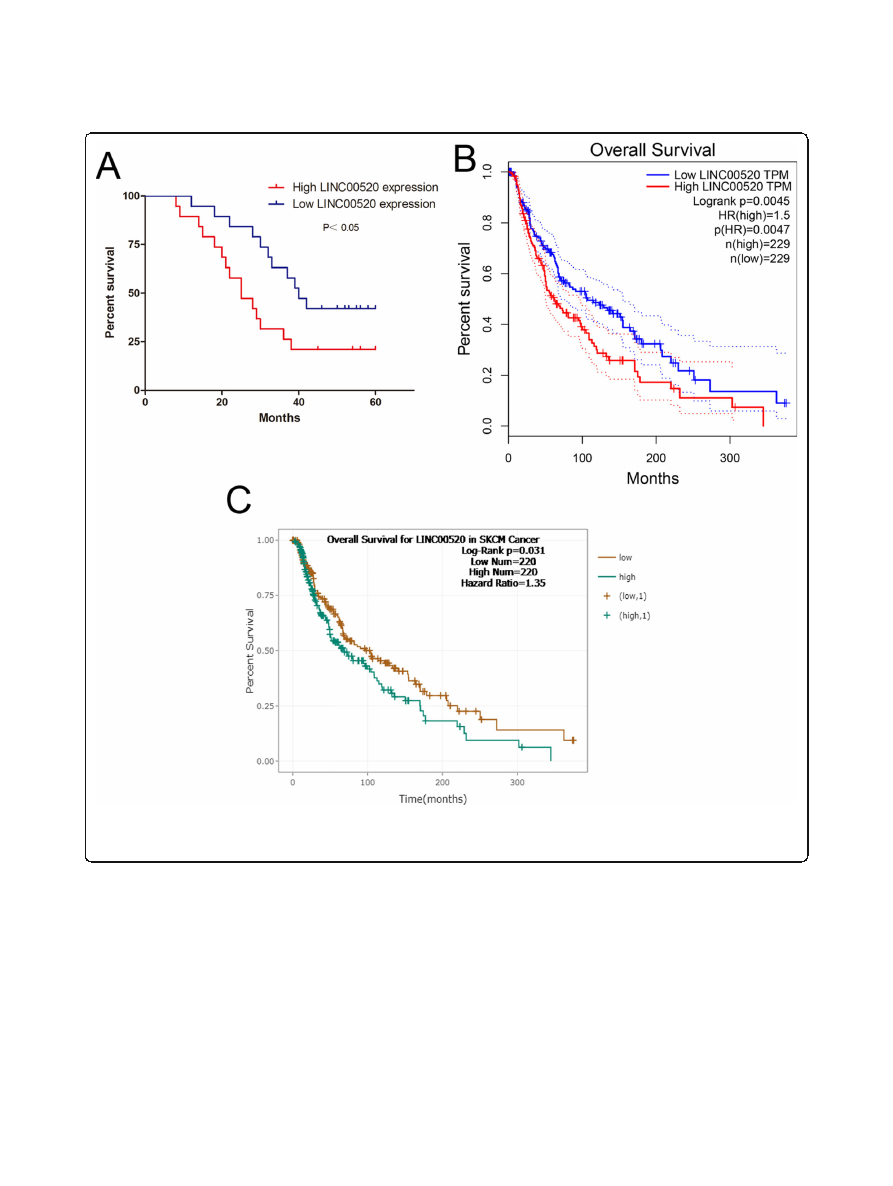
Fig. 2
LINC00520 is an risk factor for the survival of patients with melanoma.
a
The overall survival curves of 38 melanoma patients with high and
low LINC00520 levels.
b
The prognostic data of melanoma in TCGA was analyzed by using GEPIA (
c
We analyzed the
prognostic data of melanoma in TCGA by using Starbase (
). Data were expressed as the mean ± SD
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 7 of 16
LINC00520 promotes the growth and metastasis of
melanoma cell through miR-125b-5p/EIF5A2 axis
EIF5A2 has been shown to act as a new oncogene in many
tumors, including melanoma
]. We discussed
whether LINC00520 exerts its oncogenic effect in melan-
oma by regulating EIF5A2 expression. LINC00520 siRNA,
miR-125b-5p mimic, miR-125b-5p mimic together with
EIF5A2 plasmid and LINC00520 siRNA together with
miR-125b-5p inhibitor were transfected into melanoma
cells. Western blotting was used to detect the change of
EIF5A2 expression in different treatment groups (Fig.
.
The miR-125b-5p mimics inhibited the proliferation,
EMT, invasion and migration of melanoma cells, and this
effect were attenuated by the EIF5A2 plasmid (Fig.
e).
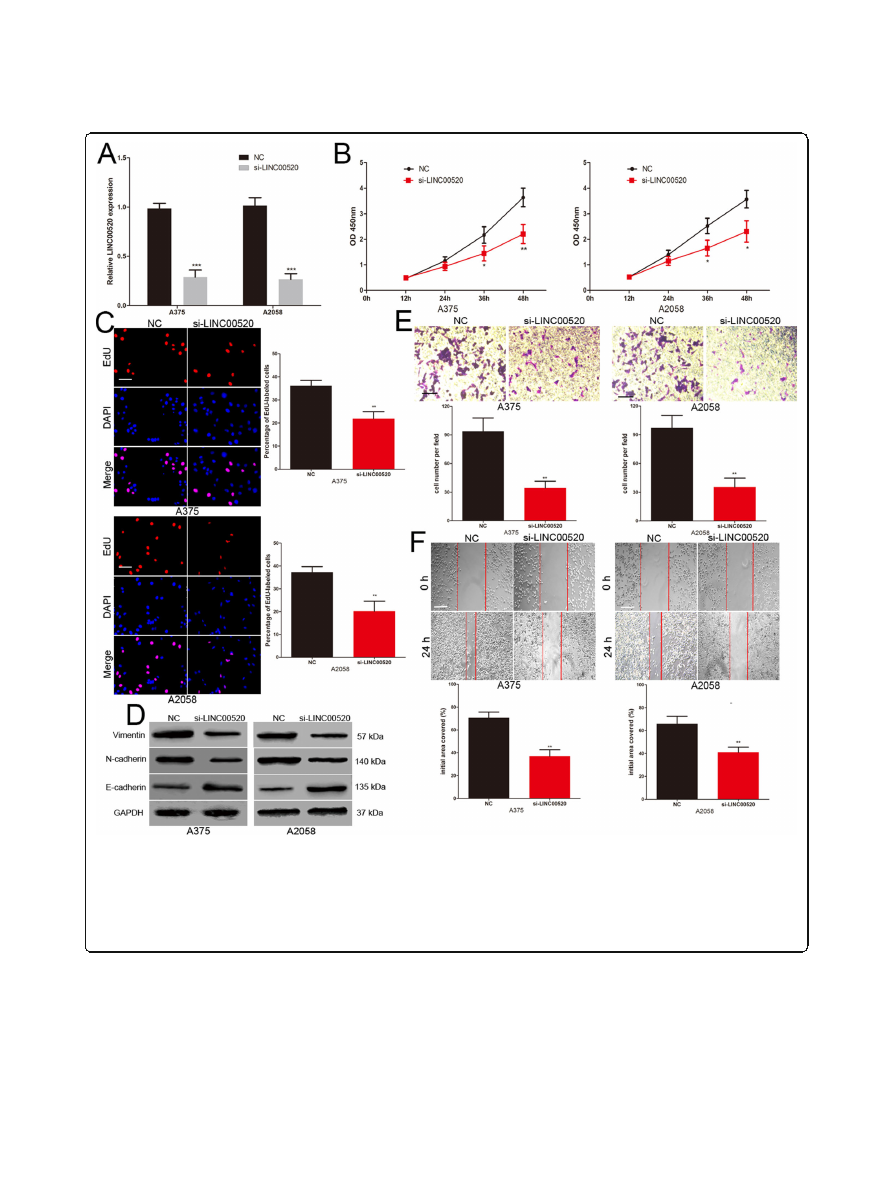
Fig. 3
LINC00520 promotes the proliferation, invasion and migration of melanoma cell.
a
Transfection efficiency of LINC00520 siRNA was
determined by PCR.
b
The proliferative ability of melanoma cells was determined by CCK8 assay in different groups.
c
The DNA synthesis of
melanoma cells grown was detected by EdU assay after transfection with NC or LINC00520 siRNA. Scale bar, 100
μ
m.
d
Western blots identified
N-cadherin, E-cadherin and Vimentin protein expression changes in NC or LINC00520 siRNA transfected melanoma cells, GAPDH was used as a
control.
e
The invasive capacity of NC or LINC00520 siRNA transfected melanoma cells was assessed by transwell assay. Scale bar, 50
μ
m.
f
Migration of melanoma cells in different transfection groups was detected by scratch wound assay. Scale bar, 100
μ
m. Data were expressed as
the mean ± SD, *P < 0.05, **
P
< 0.01, ***
P
< 0.001
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 8 of 16
These suggested that miR-125b-5p plays the role of tumor
suppressor in melanoma by targeting EIF5A2. Moreover,
we found that the inhibitory effect of LINC00520 siRNA
on the EIF5A2 expression, proliferation, EMT, invasion
and migration of melanoma cells were reversed by miR-
125b-5p inhibitor (Fig.
e). We also found that miR-
125b-5p inhibitor abolish the role of LINC00520 siRNA
on the EIF5A2 expression, proliferation and metastasis of
BRAF-WT melanoma cells (Supplementary Fig.
D).
This results further strengthen our conclusions in the
context of melanoma independently from the BRAF
mutated. Take together, these results indicated that
LINC00520 promotes the growth and metastasis of mel-
anoma by decoying miR-125b-5p to promote EIF5A2
expression.
LINC00520 exerts its pro-growth and pro-metastasis
activity through regulating miR-125b-5p/EIF5A2 axis
in vivo
Finally, we studied the effect of LINC00520 on the
growth and metastasis of melanoma cells in vivo. Stably
expressing LINC00520 shRNA or lentiviral control A375
cells were subcutaneously injected into 10 nude mice
(Fig.
a). After 28 days, the nude mice were sacrificed
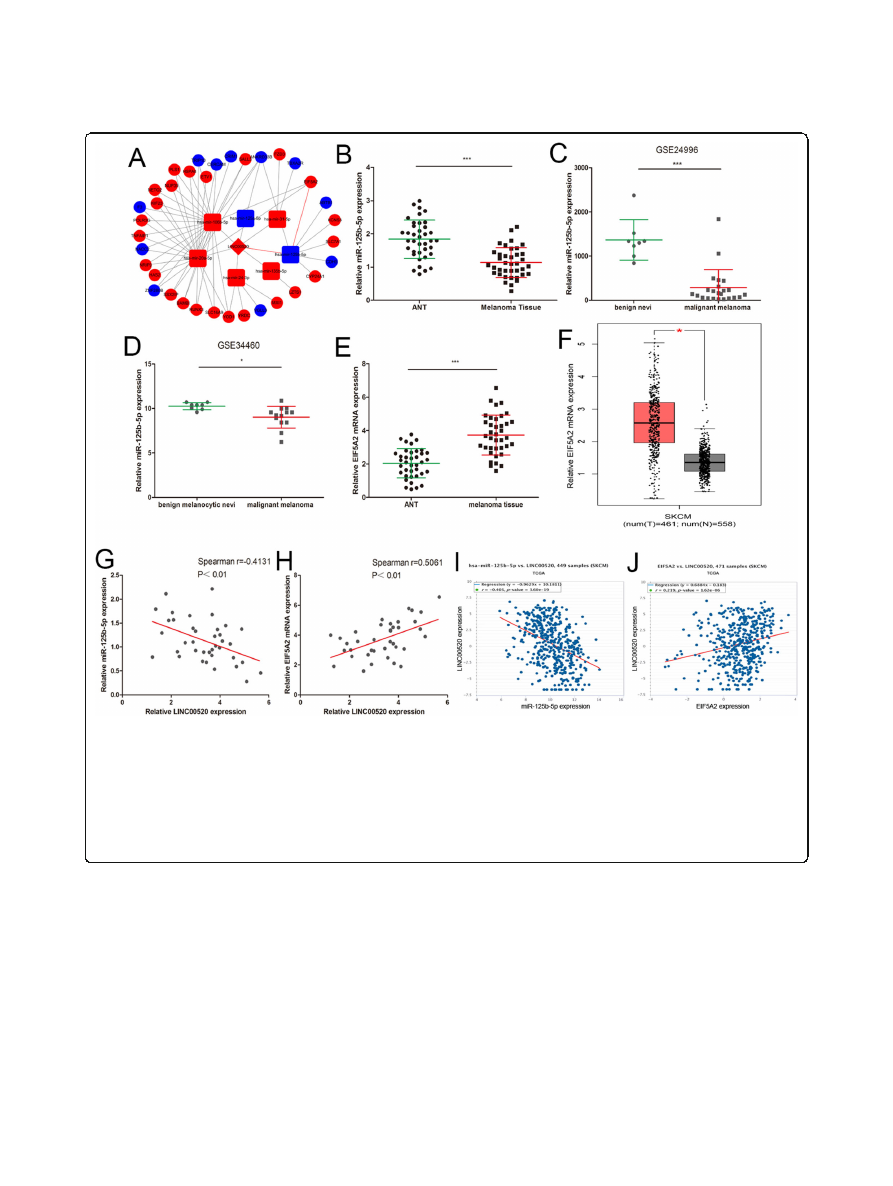
Fig. 4
ceRNA analysis of LINC00520.
a
Cytoscape was used to visualize LINC00520-miRNA-target gene nectworks based on our miRNA-seq and RNA-
seq data. The interaction between LINC00520 and miRNAs was predicted through miRcode. TargetScan, miRDB and miRTarBase were used to find the
target genes of miRNAs. Red color correspond to high expression, and blue color correspond to low expression.
b
The miR-125b-5p expression were
detected in 38 malignant melanoma tissues.
c
The expression of miR-125b-5p was analyzed by using GEO#GSE24996 dataset.
d
The expression of miR-
125b-5p was detected in GEO#GSE34460 dataset.
e
The expression of EIF5A2 was analyzed in 38 malignant melanoma tissues and ANT.
f
The
expression of EIF5A2 was detected from TCGA by using GEPIA (
).
g
The correlation of LINC00520 and miR-125b-5p in 38
melanoma tissues was negative.
h
The positive correlation between LINC00520 and EIF5A2 mRNA in 38 melanoma tissues.
i
A negative correlation
between LINC00520 and miR-125b-5p expression were found in TCGA melanoma dataset.
j
The positive correlation between LINC00520 and EIF5A2
mRNA levels in TCGA melanoma dataset. Data were expressed as the mean ± SD, *
P
< 0.05, ***P < 0.001
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 9 of 16
and the excision tumour is shown in Fig.
b. Knockdown
group of LINC00520 displayed the inhibition of tumor
growth compared to the control group between 16 and
28 days (Fig.
c). The weight of tumour in the
LINC00520 shRNA group was lighter than the control
group (Fig.
The xenograft tumour tissues were con-
firmed by H&E staining (Fig.
e), and the expression of
LINC00520, miR-125b-5p and EIF5A2 in the sections of
excision tumour were detected. The miR-125b-5p level
was increased and the EIF5A2 level was decreased with
the knockdown of LINC00520 (Fig.
f and g). To further
investigated the effects of LINC00520 on the metastasis
of melanoma cells in vivo, A375 cells that stably express-
ing LINC00520 shRNA or control were tail vein-injected
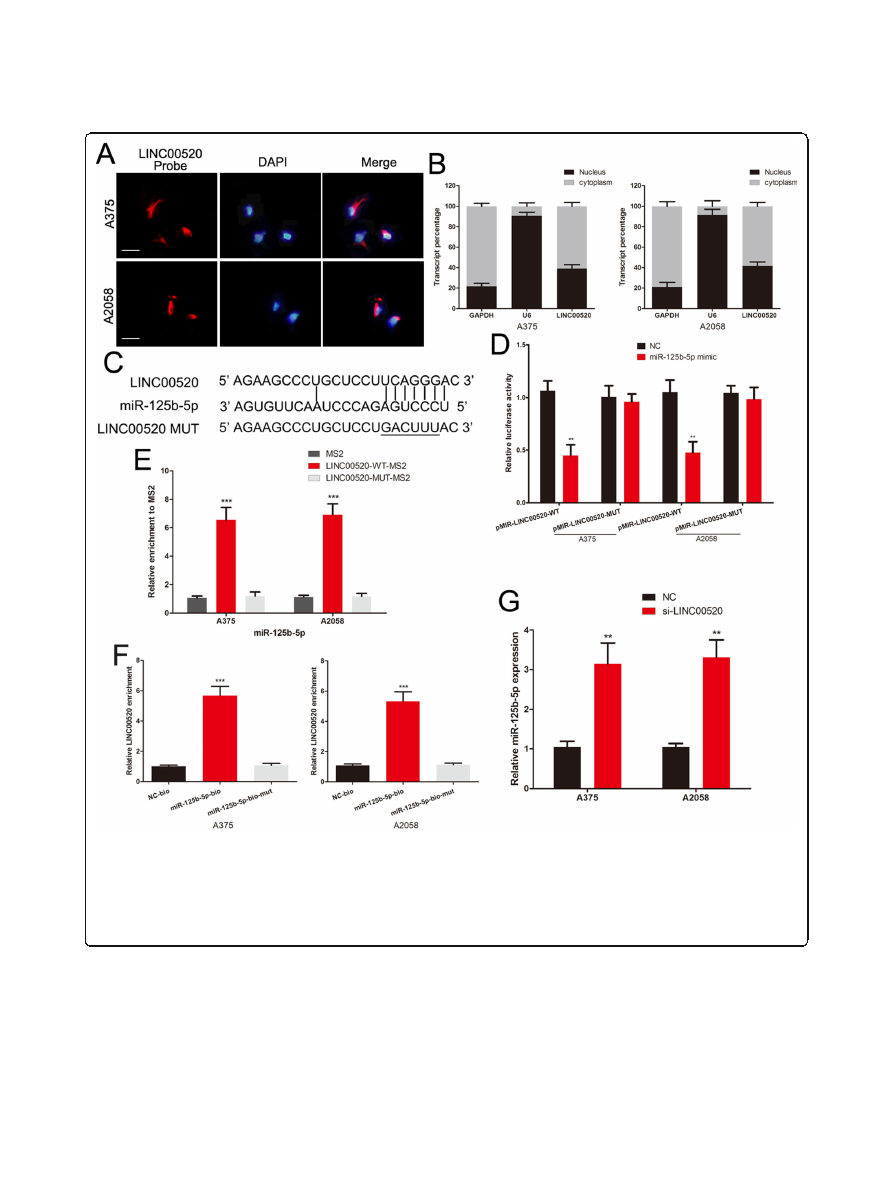
Fig. 5
LINC00520 sponges miR-125b-5p in melanoma cells.
a
FISH showed that LINC00520 was mainly distributed in both cytoplasm and nucleus
in melanoma cells. Scale bar, 25
μ
m.
b
qRT-PCR of nuclear and cytoplasm RNA fractions detected the LINC00520 expression in cytoplasm and
nuclear.
c
The binding sites of miR-125b-5p on LINC00520, and target sequences were mutated.
d
Luciferase assay of melanoma cells transfected
with LINC00520-WT or LINC00520-MUT reporter together with miR-125b-5p or NC.
e
MS2-RIP was used to detect the endogenous miR-125b-5p
associated with the MS2-tagged LINC00520.
f
Melanoma cells transfected with biotin-labeled miR-125b-5p, mutated or NC oligos, and assayed by
biotin-based pull down. The expression of LINC00520 were detected by qRT
–
PCR.
g
The levels of miR-125b-5p in melanoma cells following
transfection with LINC00520 siRNA or NC. Data were expressed as the mean ± SD, **
P
< 0.01, ***
P
< 0.001
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 10 of 16
into nude mice (Fig.
a). Knockdown group of
LINC00520 showed lower levels of lung colonisation
compared with the control group (Fig.
Silencing of
LINC00520 decreased the number of metastatic lung
nodules (Fig.
i), and HE staining confirmed the meta-
static lung tumor tissues (Fig.
j). On the sections of
metastatic pulmonary nodules, miR-125b-5p was in-
creased and EIF5A2 was reduced in the LINC00520
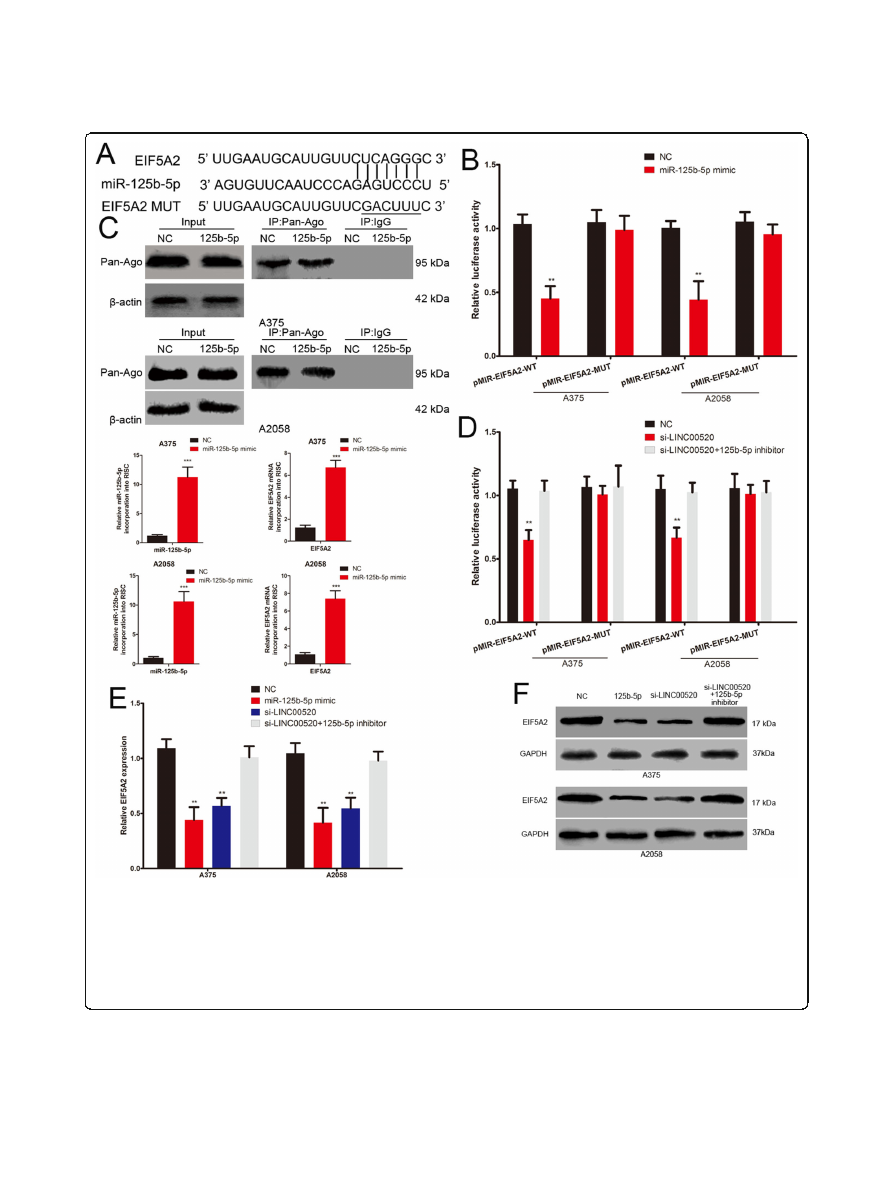
Fig. 6
LINC00520 acts as a ceRNA to promote EIF5A2 expression.
a
The binding sites of miR-125b-5p on the 3
′
-UTR of EIF5A2, and target
sequences were mutated.
b
Luciferase assay of cells transfected with EIF5A2
–
3
′
UTR-WT or EIF5A2
–
3
′
UTR-MUT reporter together with miR-125b-
5p mimic or NC.
c
Immunoprecipitation (up) of the Ago2/RISC using the Pan-Ago2 antibody in overexpressing miR-125b-5p melanoma cells. IgG
was used as a negative control and
β
-actin was used as an internal control. PCR analysis (down) of miR-125b-5p and EIF5A2 incorporated into
RISC in overexpressing miR-125b-5p melanoma cells.
d
Luciferase assay of cells transfected with EIF5A2
–
3
′
UTR-WT or EIF5A2
–
3
′
UTR-MUT reporter
together with LINC00520 siRNA or LINC00520 siRNA plus miR-125b-5p inhibitor.
e
The expression of EIF5A2 mRNA in melanoma cells transfected
with miR-125b-5p mimic, LINC00520 siRNA or LINC00520 siRNA plus miR-125b-5p inhibitor.
f
Western blots identified EIF5A2 protein expression
changes in different groups, GAPDH was used as a control. Data were expressed as the mean ± SD, **P < 0.01, ***P < 0.001
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 11 of 16
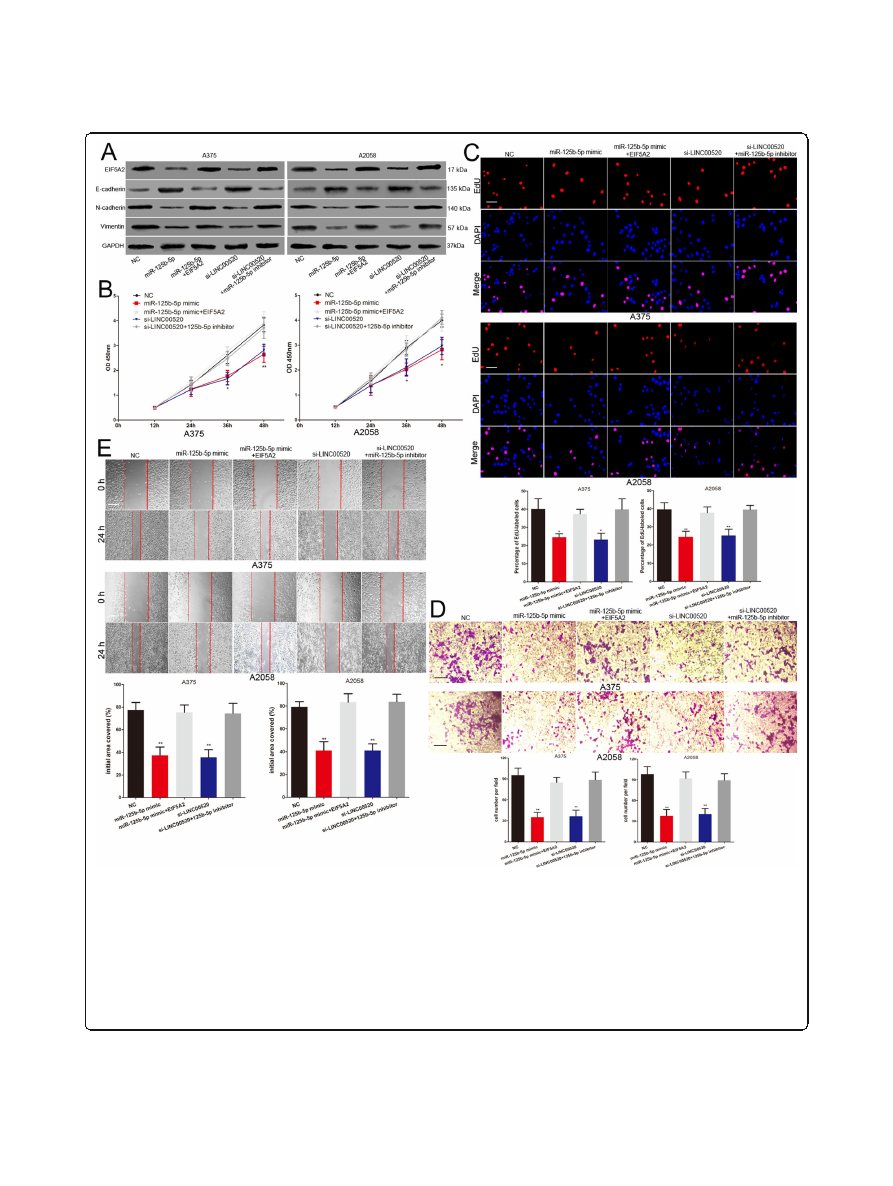
Fig. 7
LINC00520 promotes the growth and metastasis of melanoma cell through miR-125b-5p/EIF5A2 axis.
a
Western blots identified EIF5A2, N-
cadherin, E-cadherin and Vimentin protein expression changes in NC, miR-125b-5p mimic, miR-125b-5p mimic plus EIF5A2, si-LINC00520 or si-
LINC00520 plus miR-125b-5p inhibitor transfected melanoma cells, GAPDH was used as a control.
b
Effect of miR-125b-5p mimic and si-LINC00520 on
the proliferative ability of melanoma cells was determined by CCK8 assay. The results were further confirmed by co-transfection EIF5A2 plasmid and
miR-125b-5p inhibitor respectively.
c
The DNA synthesis of melanoma cells grown was detected by EdU assay following transfection with NC, miR-
125b-5p mimic, miR-125b-5p mimic plus EIF5A2, si-LINC00520 or si-LINC00520 plus miR-125b-5p inhibitor. Scale bar, 100
μ
m.
d
Effect of miR-125b-5p
mimic and si-LINC00520 on the invasive capacity of melanoma cells was assessed by transwell assay. The results were verified by the recovery
experiment of co-transfection EIF5A2 plasmid and miR-125b-5p inhibitor respectively. Scale bar, 50
μ
m.
e
Migration of melanoma cells was detected
by scratch wound assay. EIF5A2 plasmid and miR-125b-5p inhibitor reversed the effect of miR-125b-5p mimic and si-LINC00520 on the migration
capability of melanoma cells respectively. Scale bar, 100
μ
m. Data were expressed as the mean ± SD, *
P
< 0.05, **P < 0.01
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 12 of 16
knockdown group (Fig.
k and l). These results demon-
strated that LINC00520 promotes the growth and me-
tastasis of melanoma through regulating miR-125b-5p/
EIF5A2 axis (Fig.
Discussion
Recently, numerous studies have revealed that some
lncRNAs play crucial role in the development and
progression of many human tumors [
].
LINC00520, located on chromosome 14, is a novel identi-
fied lncRNA. LINC00520 has been shown to up-regulate
and modulate the malignant phenotype of tumor cells in
some malignant tumors
,
,
]. LINC00520 promotes
the proliferation, migration and invasion of glioma cells,
but inhibits its apoptosis
]. Wu et al. reported that
LINC00520 contribute to the metastasis of laryngeal
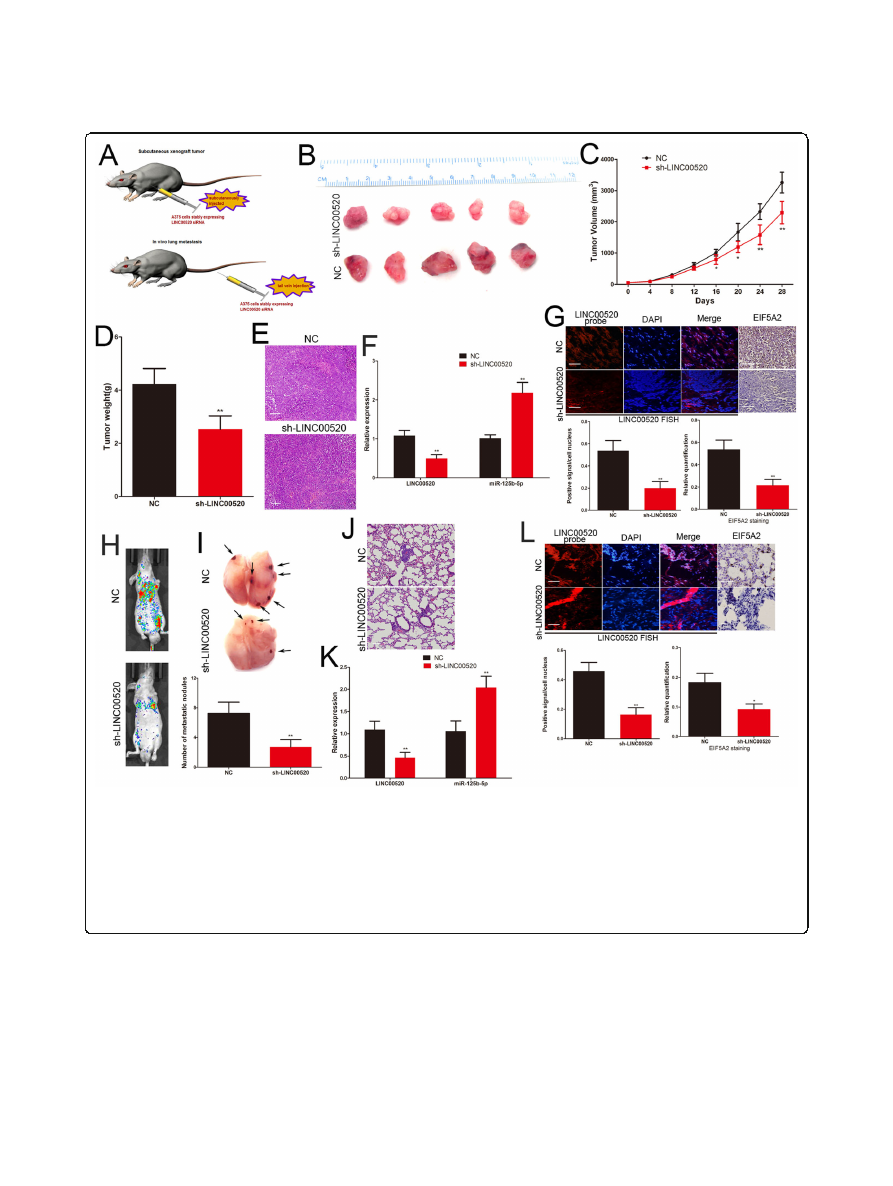
Fig. 8
LINC00520 exerts its pro-growth and pro-metastasis activity through regulating miR-125b-5p/EIF5A2 axis in vivo.
a
The schema of the animal
experiment.
b
The excision tumor in nude mice of A375 cells that stably expressing LINC00520 shRNA or control xenografts.
c
Differences in the volume of
tumor among groups.
d
The tumor weight of excised tumor tissues.
e
Xenograft tumour tissues were confirmed by H&E staining. Scale bar, 50
μ
m.
f
Expressions of LINC00520 and miR-125b-5p in xenograft tumour tissues were detect by qRT-PCR.
g
The expression of LINC00520 and EIF5A2 were
examined by FISH and immunohistochemistry of sections from the xenograft tumour tissues. Scale bar, 25
μ
m.
h
Representative bioluminescence images
of mice after tail vein injection of stably expressing LINC00520 shRNA or control A375 cells.
i
The excision lung tissues in nude mice, LINC00520 shRNA
caused a decrease in the number of metastatic lung nodules.
j
Metastatic lung nodules were confirmed by H&E staining. Scale bar, 50
μ
m.
k
qRT-PCR
detected the LINC00520 and miR-125b-5p level in the metastatic lung nodules.
l
The expression of LINC00520 and EIF5A2 were detected by FISH and
immunohistochemistry of sections from the metastatic lung nodules. Scale bar, 25
μ
m. Data were expressed as the mean ± SD, *P < 0.05, **P < 0.01
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 13 of 16
squamous cell carcinoma
. However, the role of
LINC00520 in malignant melanoma has not been studied
until now. In this study, we first analyzed the lncRNAs ex-
pression profile of melanoma tissue, and found that
LINC00520 was increased in melanoma. We verified the re-
sults of lncRNAs expression profile using more samples,
and found that high LINC00520 level conferred a poorer
prognosis to melanoma patients based on the analysis of
our samples and public database. LINC00520 has also been
demonstrated to promote the proliferation, invasion and
migration of melanoma cell.
We further explored the mechanism of LINC00520 in
melanoma. A handle of studies have been proved that
some lncRNAs can act as ceRNAs in the malignant pro-
gression of many tumors
ceRNAs reduce the
binding of miRNAs to target genes by decoying miRNAs,
thus regulating the expression of specific genes [
,
]. It
is reported that LINC00520 exhibits pro-oncogenic func-
tion in nasopharyngeal carcinoma by regulating the miR-
26b-3p/USP39 axis [
]. We subsequently constructed the
LINC00520-miRNA-target gene network based on our
miRNA-seq and RNA-seq data and bioinformatics predic-
tions. LINC00520, miR-125b-5p and EIF5A2 were found
to have a potential ceRNA correlation in melanoma. Our
melanoma samples and public database further confirmed
the network of LINC00520, miR-125b-5p and EIF5A2.
We demonstrated that LINC00520 directly binds to miR-
125b-5p by using Luciferase reporter assay, MS2-RIP
assay and RNA pull-down assay. EIF5A2 has also been
proved to be the target gene of miR-125b-5p in melan-
oma. LINC00520 siRNA reperssed the expression of
EIF5A2 and the luciferase activity of wild type EIF5A2
3
’
UTR luciferase vectors, and this repression were attenu-
ated by miR-125b-5p inhibitor. All results suggested that
LINC00520 promotes EIF5A2 expression by decoying
miR-125b-5p in melanoma.
It has been proved that miR-125b-5p acts as a tumor sup-
pressor in the malignant progress of many human tumors
[
,
In particular, miR-125b-5p is an independent pre-
dictor of survival in melanoma patients, and miR-125b-5p is
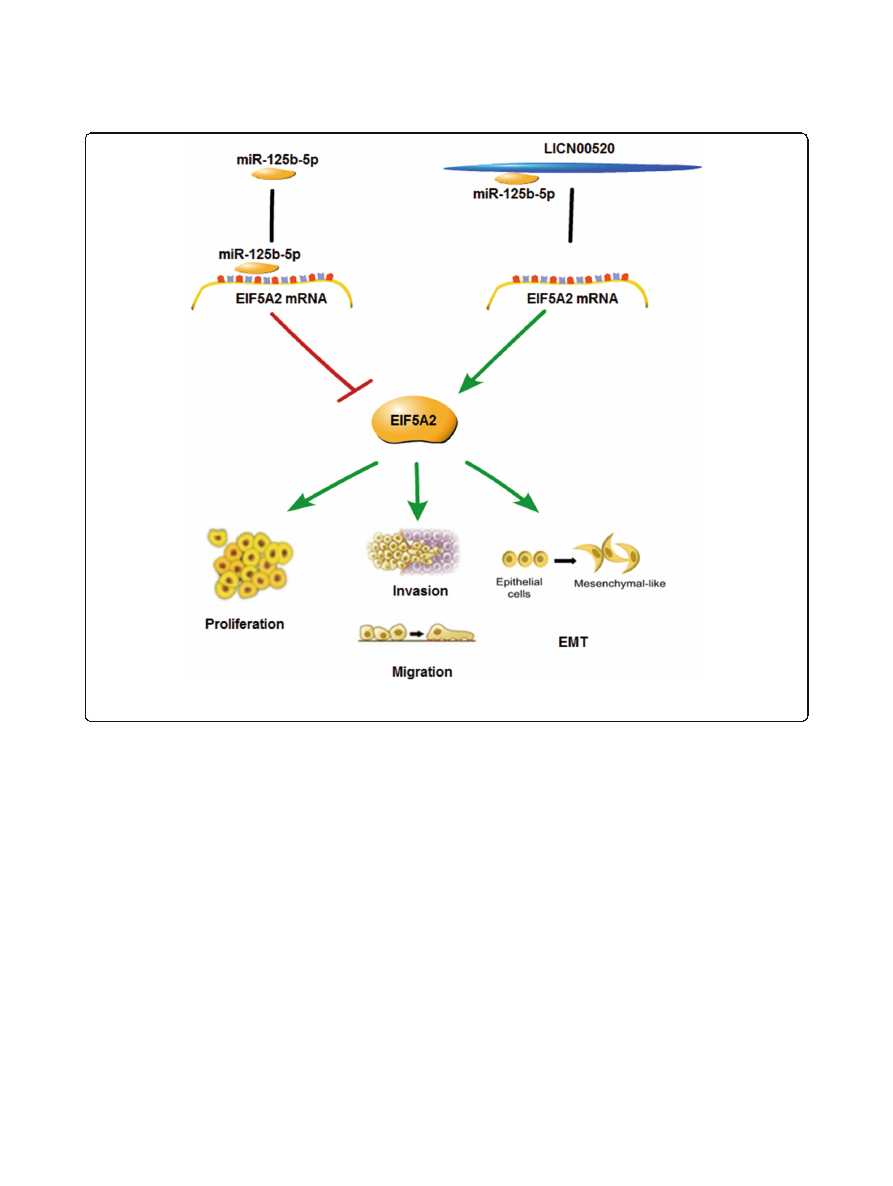
Fig. 9
A schematic model showing that LINC00520 facilitates the growth and metastasis of malignant melanoma by competitively binding to
miR-125b-5p to liberate EIF5A2 mRNA transcripts
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 14 of 16

down-regulated and suppresses the proliferation and inva-
sion of melanoma
,
]. miR-125b-5p were shown to be
involved in the vemurafenib resistance of resistant BRAF-
mutant melanoma cell
]. EIF5A2, the member of the EIF
family, is a novel oncogene and up-regulated in ovarian can-
cer, esophageal cancer, gastric cancer, etc.
].
EIF5A2 participates in many biological processes of tumor
cells, including growth, metastasis and EMT
,
,
]. It
was found that EIF5A2 also plays a role of oncogene in mel-
anoma
]. Here, we confirmed that miR-125b-5p exert its
anti-proliferation and anti-metastasis effects by targeting
EIF5A2 in melanoma. Moreover, we demonstrated that the
effect of LINC00520 siRNA on the proliferation, EMT, inva-
sion and migration of melanoma cells were reversed by
miR-125b-5p inhibitor. LINC00520 also promotes melan-
oma growth and metastasis in vivo by regulating miR-125b-
5p/EIF5A2 axis. We also domnostrated that the role of
LINC00520 in melanoma cells is independent of BRAF
mutation. Take together, our research reveal the influence
of LINC00520/miR-125b-5p/EIF5A2 on the biological pro-
gression of melanoma. Huber et al. have reported that miR-
125b-5p is released in the circulation and associated with
immunotherapy of melanoma [
Therefore, to detect the
expression level of LINC00520 in blood circulation and
explore its value in the clinical diagnosis and treatment of
patients with melanoma is our future research direction.
Conclusion
In conclusion, all results indicated that LINC00520 plays the
pivotal role in the development of melanoma. LINC00520 fa-
cilitates the growth and metastasis of malignant melanoma
by competitively binding to miR-125b-5p to liberate EIF5A2
mRNA transcripts, thereby promotes the EIF5A2 expression.
Understanding the molecular mechanism of LINC00520 in
melanoma is important to improve our knowledge of the
molecular biological of malignant progression of melanoma.
The deep study of LINC00520/miR-125b-5p/EIF5A2 axis is
helpful for us to identify new biomarkers or therapeutic tar-
get for melanoma patients.
Supplementary information
Supplementary information
accompanies this paper at
.
Additional file 1: Figure S1.
(A) Western blots identified EIF5A2 protein
expression changes in NC, si-LINC00520 or si-LINC00520 plus miR-125b-
5p inhibitor transfected MeWo cells, GAPDH was used as a control. (B) Ef-
fect of si-LINC00520 on the proliferative ability of MeWo cells was deter-
mined by CCK8 assay, and the results were further confirmed by co-
transfection miR-125b-5p inhibitor. (C) The invasive capacity of MeWo
cells was detected by transwell assay following transfection with NC, si-
LINC00520 or si-LINC00520 plus miR-125b-5p inhibitor. (D) The migratory
ability of MeWo cells was assessed by the scratch wound assay. miR-
125b-5p inhibitor reversed the effect of si-LINC00520 on the migration
capability of MeWo cells. Scale bar, 100
μ
m. Data were expressed as the
mean ± SD, *
P
< 0.05, **
P
< 0.01, ***
P
< 0.001.
Abbreviations
LINC00520:
Long intergenic non-protein coding RNA520; EIF5A2: Eukaryotic
initiation factor 5A2; LncRNAs: Long non-coding RNAs; CCK-8: Cell counting
kit-8; EMT: Epithelial-to-mesenchymal transition; TCGA: The Cancer Genome
Atlas; DMEM: Dulbecco
’
s modified Eagle
’
s medium; ATCC: American Type
Culture Collection; shRNA: short hairpin RNA; siRNA: Small interfering RNA;
PI: Propidium iodide; RISC: RNA-induced silencing complex;
FISH: Fluorescence in situ hybridization
Acknowledgements
Not applicable.
Authors
’
contributions
WKL and XFB conceived and designed the experiments. YTD, HTY, SJM, HRR,
JLW and FL performed the experiments. WKL and XFB provided the
technical support. WKL, HTY, YTD and XFB analyzed and interpreted the data.
WKL, HTY and YTD wrote the manuscript. The author(s) read and approved
the final manuscript.
Funding
This study was funded by National Natural Science Foundation of China
(81802726).
Availability of data and materials
All the data and materials supporting the conclusions were included in the
main paper.
Ethics approval and consent to participate
The study was conducted in accordance with the Declaration of Helsinki
principles. It was approved by the Ethics Committee of the Affiliated People
’
s
Hospital of Jiangsu University.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Author details
1
Department of Plastic Surgery, Affiliated People
’
s Hospital of Jiangsu
University, 8 Dianli Road, Zhenjiang 212000, Jiangsu, China.
2
Department of
Rehabilitation, Changshu No. 2 People
’
s Hospital (The 5th Clinical Medical
College of Yangzhou University), Changshu, Jiangsu, China.
3
Department of
General Surgery, Affiliated People
’
s Hospital of Jiangsu University, 8 Dianli
Road, Zhenjiang 212000, Jiangsu, China.
Received: 13 February 2020 Accepted: 19 May 2020
References
1.
Haass NK, Schumacher U. Melanoma never says die. Exp Dermatol. 2014;
23(7):471
–
2.
2.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;
67(1):7
–
30.
.
3.
Kosnopfel C, Sinnberg T, Sauer B, Busch C, Niessner H, Schmitt A, et al. YB-1
expression and phosphorylation regulate Tumorigenicity and invasiveness
in melanoma by influencing EMT. Mol Cancer Res. 2018;16(7):1149
–
60.
4.
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer statistics
in China, 2015. CA Cancer J Clin. 2016;66(2):115
–
32.
5.
Tsao H, Chin L, Garraway LA, Fisher DE. Melanoma: from mutations to
medicine. Genes Dev. 2012;26(11):1131
–
55.
.
6.
Paluncic J, Kovacevic Z, Jansson PJ, Kalinowski D, Merlot AM, Huang ML,
et al. Roads to melanoma: key pathways and emerging players in
melanoma progression and oncogenic signaling. Biochim Biophys Acta.
2016;1863(4):770
–
84.
7.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol
Cell. 2011;43(6):904
–
14.
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 15 of 16

8.
Schmitt AM, Chang HY. Long noncoding RNAs in Cancer pathways. Cancer
Cell. 2016;29(4):452
–
63.
.
9.
Lingadahalli S, Jadhao S, Sung YY, Chen M, Hu L, Chen X, et al. Novel
lncRNA LINC00844 regulates prostate Cancer cell migration and invasion
through AR signaling. Molecular cancer research : MCR. 2018;16(12):1865
–
78.
10.
Luan W, Ding Y, Ma S, Ruan H, Wang J, Lu F. Long noncoding RNA
LINC00518 acts as a competing endogenous RNA to promote the
metastasis of malignant melanoma via miR-204-5p/AP1S2 axis. Cell Death
Dis. 2019;10(11):855.
11.
Luan W, Zhang X, Ruan H, Wang J, Bu X. Long noncoding RNA OIP5-AS1
acts AS a competing endogenous RNA to promote glutamine catabolism
and malignant melanoma growth by sponging miR-217. J Cell Physiol.
2019.
12.
Henry WS, Hendrickson DG, Beca F, Glass B, Lindahl-Allen M, He L, et al.
LINC00520 is induced by Src, STAT3, and PI3K and plays a functional role in
breast cancer. Oncotarget. 2016;7(50):81981
–
94.
13.
Wu YY, Gao W, Zhang YL, Niu M, Cui JJ, Xiang CX, et al. Expression and
clinical significance of long non-coding RNA LINC00520 in laryngeal
squamous cell carcinoma. Lin chuang er bi yan hou tou jing wai ke za zhi.
2018;32(2):91
–
5.
.
14.
Xie T, Pi G, Yang B, Ren H, Yu J, Ren Q, et al. Long non-coding RNA
520 is a negative prognostic biomarker and exhibits pro-oncogenic
function in nasopharyngeal carcinoma carcinogenesis through
regulation of miR-26b-3p/USP39 axis. Gene. 2019;707:44
–
52.
.
15.
Mei XL, Zhong S. Long noncoding RNA LINC00520 prevents the
progression of cutaneous squamous cell carcinoma through the
inactivation of the PI3K/Akt signaling pathway by downregulating EGFR.
Chin Med J. 2019;132(4):454
–
65.
.
16.
Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and
competition. Nature. 2014;505(7483):344
–
52.
17.
Luan W, Li L, Shi Y, Bu X, Xia Y, Wang J, et al. Long non-coding RNA
MALAT1 acts as a competing endogenous RNA to promote malignant
melanoma growth and metastasis by sponging miR-22. Oncotarget. 2016;
7(39):63901
–
12.
18.
Jenkins ZA, Haag PG, Johansson HE. Human eIF5A2 on chromosome 3q25-
q27 is a phylogenetically conserved vertebrate variant of eukaryotic
translation initiation factor 5A with tissue-specific expression. Genomics.
2001;71(1):101
–
9.
19.
Bao Y, Lu Y, Wang X, Feng W, Sun X, Guo H, et al. Eukaryotic translation
initiation factor 5A2 (eIF5A2) regulates chemoresistance in colorectal cancer
through epithelial mesenchymal transition. Cancer Cell Int. 2015;15:109.
20.
Liu RR, Lv YS, Tang YX, Wang YF, Chen XL, Zheng XX, et al. Eukaryotic
translation initiation factor 5A2 regulates the migration and invasion of
hepatocellular carcinoma cells via pathways involving reactive oxygen
species. Oncotarget. 2016;7(17):24348
–
60.
21.
Sun J, Xu Z, Lv H, Wang Y, Wang L, Ni Y, et al. eIF5A2 regulates the
resistance of gastric cancer cells to cisplatin via induction of EMT. Am J
Transl Res. 2018;10(12):4269
–
79.
22.
Li Y, Fu L, Li JB, Qin Y, Zeng TT, Zhou J, et al. Increased expression of
EIF5A2, via hypoxia or gene amplification, contributes to metastasis and
angiogenesis of esophageal squamous cell carcinoma. Gastroenterology.
2014;146(7):1701
–
13 e9.
23.
Khosravi S, Wong RP, Ardekani GS, Zhang G, Martinka M, Ong CJ, et al. Role of
EIF5A2, a downstream target of Akt, in promoting melanoma cell invasion. Br J
Cancer. 2014;110(2):399
–
408.
24.
Ni X, Ding Y, Yuan H, Shao J, Yan Y, Guo R, et al. Long non-coding RNA
ZEB1-AS1 promotes colon adenocarcinoma malignant progression via miR-
455-3p/PAK2 axis. Cell Proliferat. 2020;53(1):e12723.
25.
Liu H, Dai C, Wu Q, Liu H, Li F. Expression profiling of long noncoding RNA
identifies lnc-MMP3-1 as a prognostic biomarker in external auditory canal
squamous cell carcinoma. Cancer medicine. 2017;6(11):2541
–
51.
.
26.
Subramanian M, Li XL, Hara T, Lal A. A biochemical approach to identify
direct microRNA targets. Methods Mol Biol. 2015;1206:29
–
37.
27.
Luan W, Wang Y, Chen X, Shi Y, Wang J, Zhang J, et al. PKM2 promotes
glucose metabolism and cell growth in gliomas through a mechanism
involving a let-7a/c-Myc/hnRNPA1 feedback loop. Oncotarget. 2015;6(15):
13006
–
18.
28.
Xu Y, Qin L, Sun T, Wu H, He T, Yang Z, et al. Twist1 promotes breast cancer
invasion and metastasis by silencing Foxa1 expression. Oncogene. 2017;
36(8):1157
–
66.
29.
Davis FM, Azimi I, Faville RA, Peters AA, Jalink K, Putney JW Jr, et al.
Induction of epithelial-mesenchymal transition (EMT) in breast cancer cells
is calcium signal dependent. Oncogene. 2014;33(18):2307
–
16.
.
30.
Li Y, Wang L, Rivera-Serrano EE, Chen X, Lemon SM. TNRC6 proteins
modulate hepatitis C virus replication by spatially regulating the binding of
miR-122/Ago2 complexes to viral RNA. Nucleic Acids Res. 2019;47(12):6411
–
24.
31.
Luan W, Li R, Liu L, Ni X, Shi Y, Xia Y, et al. Long non-coding RNA HOTAIR
acts as a competing endogenous RNA to promote malignant melanoma
progression by sponging miR-152-3p. Oncotarget. 2017;8(49):85401
–
14.
32.
Davalos V, Esteller M. Disruption of Long noncoding RNAs targets Cancer
Hallmark pathways in lung tumorigenesis. Cancer Res. 2019;79(12):3028
–
30.
.
33.
Chen B, Wang C, Zhang J, Zhou Y, Hu W, Guo T. New insights into long
noncoding RNAs and pseudogenes in prognosis of renal cell carcinoma.
Cancer Cell Int. 2018;18:157.
34.
Wang Y, Yang C, Liu X, Zheng J, Zhang F, Wang D, et al. Transcription factor
AP-4 (TFAP4)-upstream ORF coding 66 aa inhibits the malignant behaviors
of glioma cells by suppressing the TFAP4/long noncoding RNA 00520/
microRNA-520f-3p feedback loop. Cancer Sci. 2020;111(3):891
–
906.
35.
Liu L, Shi Y, Shi J, Wang H, Sheng Y, Jiang Q, et al. The long non-coding
RNA SNHG1 promotes glioma progression by competitively binding to miR-
194 to regulate PHLDA1 expression. Cell Death Dis. 2019;10(6):463.
36.
Lu G, Li Y, Ma Y, Lu J, Chen Y, Jiang Q, et al. Long noncoding RNA LINC00511
contributes to breast cancer tumourigenesis and stemness by inducing the
miR-185-3p/E2F1/Nanog axis. J Experiment Clin Cancer Res. 2018;37(1):289.
37.
Sang M, Meng L, Liu S, Ding P, Chang S, Ju Y, et al. Circular RNA ciRS-7 maintains
metastatic phenotypes as a ceRNA of miR-1299 to target MMPs. Mol Cancer Res.
2018;16(11):1665
–
75.
.
38.
Liu S, Chen Q, Wang Y. MiR-125b-5p suppresses the bladder cancer
progression via targeting HK2 and suppressing PI3K/AKT pathway. Hum Cell.
2020;33(1):185
–
94.
39.
Hua S, Quan Y, Zhan M, Liao H, Li Y, Lu L. miR-125b-5p inhibits cell
proliferation, migration, and invasion in hepatocellular carcinoma via
targeting TXNRD1. Cancer Cell Int. 2019;19:203.
40.
Yan J, Jiang Q, Lu H, Na S, Long S, Xin Y, et al. Association between
microRNA-125b expression in formalin-fixed paraffin-embedded tumor
tissues and prognosis in patients with melanoma. Oncol Lett. 2019;18(2):
1856
–
62.
41.
Zhang J, Lu L, Xiong Y, Qin W, Zhang Y, Qian Y, et al. MLK3 promotes
melanoma proliferation and invasion and is a target of microRNA-125b. Clin
Exp Dermatol. 2014;39(3):376
–
84.
.
42.
Vergani E, Di Guardo L, Dugo M, Rigoletto S, Tragni G, Ruggeri R, et al.
Overcoming melanoma resistance to vemurafenib by targeting CCL2-
induced miR-34a, miR-100 and miR-125b. Oncotarget. 2016;7(4):4428
–
41.
43.
Bussard KM, Mutkus L, Stumpf K, Gomez-Manzano C, Marini FC. Tumor-
associated stromal cells as key contributors to the tumor microenvironment.
Breast cancer research : BCR. 2016;18(1):84.
44.
Guan XY, Fung JM, Ma NF, Lau SH, Tai LS, Xie D, et al. Oncogenic role of
eIF-5A2 in the development of ovarian cancer. Cancer Res. 2004;64(12):
4197
–
200.
.
45.
Meng QB, Kang WM, Yu JC, Liu YQ, Ma ZQ, Zhou L, et al. Overexpression of
eukaryotic translation initiation factor 5A2 (EIF5A2) correlates with cell
aggressiveness and poor survival in gastric cancer. PLoS One. 2015;10(3):
e0119229.
.
46.
Huber V, Vallacchi V, Fleming V, Hu X, Cova A, Dugo M, et al. Tumor-derived
microRNAs induce myeloid suppressor cells and predict immunotherapy
resistance in melanoma. J Clin Invest. 2018;128(12):5505
–
16.
.
Publisher
’
s Note
Springer Nature remains neutral with regard to jurisdictional claims in
published maps and institutional affiliations.
Luan
et al. Journal of Experimental & Clinical Cancer Research
(2020) 39:96
Page 16 of 16















